TL;DR
- In the context of vector search, an Inverted File List (IVF) is an index powered by K-Means clustering. The data stored in the index is partitioned or divided into
nlist clusters. In the end, the IVF tells us the centroids of the nlist clusters and which cluster each data “point” belongs to.
- Product Quantization (PQ) is a lossy compression technique (not a search technique per se) which compresses a vector by subdividing it into
M parts and quantizing each part independently using K-Means clustering once again.
- A product quantizer will expose a pair of
encode/decode methods. encode(x) will take a vector x and encode it into a compressed representation y (also known as codeword). decode(y) will output quantized vector z. The difference between z and x is the quantization error.
The Faiss IndexIVFPQ is a popular algorithm for ANN (approximate nearest neighbor) search of high-dimensional vectors.
From Faiss wiki page:
The IndexIVFPQ is probably the most useful indexing structure for large-scale search.
In this post we try to understand its internals and how it works. I also cover several gotchas (mistakes to avoid).
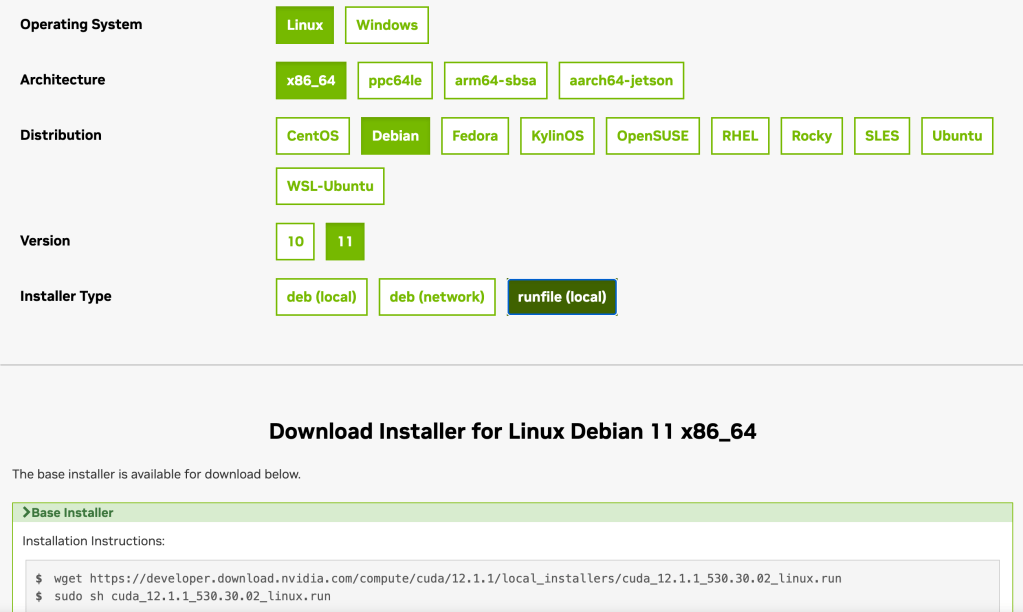
Its example usage is as follows (from the same wiki page):
coarse_quantizer = faiss.IndexFlatL2 (d)
index = faiss.IndexIVFPQ (coarse_quantizer, d,
ncentroids, code_size, 8) # Bug here! see my post later
index.nprobe = 5
Another way to create this index is using the index factory (refer the table on same wiki page. In that table they also refer to this
index as IVFADC (coarse quantizer+PQ on residuals) under the Method column):
index_type = f"IVF{nlist},PQ{m}"
faiss_ivf_pq = faiss.index_factory(d, index_type)
faiss_ivf_pq.nprobe = nprobe
When we create an IndexIVFPQ and try to perform a ANN search to get k-ANN of a given query vector,
there are two steps that occur which are explained below:
Step 1 – Find the cluster(s) in which the query vector lies. Done using a IndexIVF index.
First, the algorithm tries to find the cluster (or clusters) in which the query vector lies. The parameter nprobe
is used to control how many cluster(s) you want to search. The higher the nprobe, the better results you will get but of course
it will take more time. The upshot of this observation is that
before you can perform a search, you need to divide or partition the data into clusters (one of the many disadvantages of IndexIVFPQ algorithm; another disadvantage is that you have to declare the number of clusters in advance; this is controlled by the ncentroids parameter in the code snippet above and also written as nlist at other places in Faiss documentation).
The partitioning of the data is done using the coarse_quantizer in above. The coarse_quantizer is used to create a IndexIVF
which is nothing but the k-means algorithm in disguise.
A survey done in 2006 voted k-means amongst the top 10 algorithms in data-mining.
I completely agree esp. if you consider IndexIVFPQ which is nothing but k-means all the way down (spoiler). From the same
wiki page under the section
titled Cell-probe methods (IndexIVF* indexes):
A typical way to speed-up the process at the cost of loosing the guarantee to find the nearest neighbor is to employ a partitioning technique such as k-means
The output of k-means algorithm is the centroids of the clusters. To find the cluster(s) in which the query vector lies, we just perform
a brute force search and compute the distances of the query vector to each of the centroids and pick the nprobe closest centroids.
From the same wiki page:
Those elements are just scanned sequentially, and the search function returns the top-k (nprobe in our case) smallest distances seen so far.
To summarize:
An IndexIVF is a data structure that clusters (training) data into nlist clusters using the k-means algorithm and when it receives a search request, it returns the closest nprobe cluster(s) to the search query by performing a brute-force search against the centroids. Runtime of search = O(nlist * log(nprobe)) if implemented using a heap.
This runtime does not factor in dimensionality d of the vectors. It is assumed the time to compute distance between vectors is independent of d
(e.g., using SIMD). If SIMD or other vectorization is not available the runtime will be O(d * nlist * log(nprobe)).
Step 2 – Perform brute force search of data in those clusters. Done using a IndexPQ index.
Once we have identified the cluster(s) to search from previous step, we just perform a brute force search of the data in those clusters
to find the k-ANN. So again, we have a brute-force search. Nothing fancy. We could store the vectors in raw form in each of the clusters
and indeed we should do so if memory is not a concern. That would correspond to a IVF{nlist},Flat (or IndexIVFFlat) index
(note that IndexIVFFlat is not same as IndexIVF). From the docs
page of IndexIVFFlat:
Inverted file with stored vectors. Here the inverted file pre-selects the vectors to be searched, but they are not otherwise encoded, the code array just contains the raw float entries.
What IndexIVFPQ does instead is that it does not store the raw vectors in each cluster. Instead it stores compressed vectors
and this is where Product Quantization comes in. So what is it? (Hint: k-means coming up again) Simply this:
- Divide each vector into
M subvectors. A commonly used value of M is d/2 in which case each subvector has length l = d/M = 2
and is thus a vector in 2D space. d is length of the vectors in your dataset e.g., 128 for the SIFT image descriptors.
let’s do an example with d=10, M=5. given a vector:
x = [0.77759472, 0.28507821, 0.58818803, 0.07639025, 0.90228833,
0.13530444, 0.35468633, 0.76250536, 0.83748744, 0.08922487]
it is broken into M=5 subvectors:
[[0.77759472, 0.28507821], [0.58818803, 0.07639025], [0.90228833, 0.13530444], [0.35468633, 0.76250536], [0.83748744, 0.08922487]]
- Perform k-means clustering again independently in each of the
M subspaces. From the original paper that introduced product quantization:
The idea is to decomposes the space into a Cartesian product of low dimensional subspaces and to quantize each subspace separately.
The Lloyd quantizer, which corresponds to the kmeans clustering algorithm, finds a near-optimal codebook by iteratively assigning the vectors of a training set to centroids and re-estimating these centroids from the assigned vectors.
Note that to do this clustering an implementation might very well instantiate an IndexIVF for each of the M subspaces – this is where it gets a little mind-bending.
The number of clusters is controlled by the nbits parameter in this case. The default value of nbits is 8 (also in the code snippet
at beginning of the post) and the number of clusters
is equal to 2^nbits e.g., with nbits=8 we have 256 clusters.
Once we have done the k-means clustering, we encode each vector to be stored in the index as follows:
- Divide each vector into
M subvectors as before.
- For each subvector, find the cluster it belongs to (can be done using IndexIVF search) and store the cluster id (this is also known as the codeword).
M subcodewords will be concatenated to give the final codeword (identifier) of the vector.
Example with d=16, M=8, nbits=8:
>>> pq = ProductQuantizer(d, M, nbits)
>>> x_train=np.random.rand(N,d)
>>> pq.train(x_train)
>>> x = np.random.rand(1,d)[0] # sample vector
>>> x
array([0.19550796, 0.73512159, 0.82772364, 0.03929587, 0.99528502,
0.6065332 , 0.2902288 , 0.23997098, 0.7986942 , 0.07180022,
0.90386235, 0.33337289, 0.64734347, 0.36640519, 0.84930454,
0.76707514])
>>> c=pq.encode(x) # codeword. the codeword compresses d * 4 = 64 bytes into 8 bytes (M * nbits / 8 in general).
>>> c
BitArray('0xa78a5c5f84e91bb5')
>>> y = pq.decode(c) # quantized vector
>>> y
array([0.22070472, 0.73106791, 0.84465414, 0.03585924, 0.96638963,
0.58450194, 0.30623678, 0.22202215, 0.77308882, 0.08511855,
0.89131492, 0.35034598, 0.63585688, 0.34263507, 0.82575109,
0.76019279])
>>> np.linalg.norm(x-y) # quantization error
0.07366224496403329
Exercise: implement the ProductQuantizer yourself.
The cluster id takes up nbits. We have M subvectors. So total storage is equal to nbits * M bits. The raw vector would take
d * 4 * 8 bits since we have to store d floating point numbers and each floating point number takes up 4 bytes. This gives
us a compression ratio of (d * 4 * 8) / (nbits * M). With M = d / 2 and nbits = 8 which is a popular setting,
we get 8x compression. Another setting you could experiment with is M = d and nbits = 8 in which case we are simply
quantizing each floating point number in a vector into 256 “gray levels” (I mention it in quotes as the levels need not be equi-spaced
and will be determined by the k-means algorithm; for those who missed it, I am making an analogy to how a colored image is quantized into a
gray scale image).
To summarize:
An IndexPQ is a data structure that quantizes the data stored in it according to the PQ algorithm and performs a brute force search in response to a query. The quantization in turn relies on k-means and can be implemented using IndexIVF.
Optimizations (the fine-print)
Precomputing and caching distances between codewords
Above covers the essentials of product quantization. There is another optimization that is commonly applied.
Consider how to compute the distance between a query vector and one of the data “points” (I use quotes because a data point is really a vector):
- First, quantize (encode) the query vector the same way as all the data “points” are encoded. This is not necessary as I explain later.
- Remember, after encoding all the data “points” you no longer have access to their actual values. You only know the codes (
M ids) of each
data “point” and each code (which is nothing but id of centroid output by the k-means algorithm) is associated to a floating point centroid.
- This means, we need to determine distance between two encoded vectors. Since the number of codes is finite,
we can precompute a lookup table of distances to gain some efficiency in performance at expense of memory. This lookup table is of size
2^nbits x 2^nbits and we need to store M lookup tables for the M subspaces.
I don’t know what Faiss does but I don’t recommend storing the lookup table if nbits > 8. The performance gains will be negated by
sharp increase in memory consumption. It is better in this case to compute the distances “on-demand” every time instead of using a cache.
Another alternative to consider is that note that we don’t necessarily need to encode the query vector (Step 1 in above) to compute its distance
from another vector in the index. All we need is to decode the stored vectors. Indeed this is what is done most commonly and goes by the
name of asymmetric distance computation (ADC). It is asymmetric because in the distance computation, the query vector is not quantized
whereas the vectors stored in the index are. In this case the lookup table does not apply. If you encode the query vector as well (Step 1 above), we get
symmetric distance computation. The original PQ paper advocates ADC. It says “An asymmetric version increases precision, as it computes the
approximate distance between a vector and a code.”
Store the residual vector in each of the clusters instead of original vector
There is one other optimization I skipped to make it simple to understand. In practice, after IndexIVF has been trained,
we are storing centroids (in full floating point precision) of the nlist clusters. When we have to store a vector x in a cluster
(remember this is where PQ kicks in), its better to encode the residual of x instead of the original vector. The residual is simply the difference
of the original vector x with the centroid of the cluster in which its stored i.e., residual = x - centroid. This will improve
accuracy of the ANN search.
In a nutshell, this is what IndexIVFPQ is about.
The original paper that introduced it is again
this (recommended reading; definitely a seminal paper in CS;
I find it hard to read research papers as they contain no examples, just algebra but hopefully you will understand it after reading this post).
You must have seen following code in Faiss documentation:
index_ivfpq.train(x_train)
This will call the train method in both IndexIVF and IndexPQ and run the k-means algorithm for both. Remember that calling train
only trains the k-means algorithm. It does not store any data in the index. If you also want to store the data in the index you should
follow up train with a call to add.
Runtime Analysis
Let’s analyze the runtime of a search request by IndexPQ:
Step 1: What is the time taken to encode the query vector? To do this we have to perform M IndexIVF searches.
Recall an IndexIVF search takes O(nlist*log(nprobe)) except that in this case nlist = 2^nbits and nprobe = 1. So we get runtime of
O(M * 2^nbits).
Step 2: Now what is the time to compute distance between two codewords? Assuming we have a lookup table we can perform O(1) lookups and we need to
perform M of these for each of the subvectors. This gives runtime of O(M) or O(1) if SIMD vectorization is available.
Step 3: We can use a heap again to find k closest “points”. This is done in O(N * log(k)) time where N is now the number of vectors (data “points”) stored in IndexPQ, but it hides the cost of computing the distance
between a pair of codewords which we have seen is O(M). So combining step 2 and 3 we have O(N * log(k) * M).
Step 4: we add O(M * 2^nbits) + O(N * log(k) * M) = O(M * 2^nbits + N * log(k) * M) to give final result.
We are not done yet. To calculate the runtime of IndexIVFPQ we need to multiply runtime of IndexIVF in Step 1 with runtime of IndexPQ in Step 2.
This gives the overall runtime as:
where I have ignored SIMD optimizations. In above the $\frac{N}{nlist}$ factor is assuming the data is equally divided into the nlist clusters (asymptotic approximation).
Gotchas
Incorrect Documentation
From Faiss wiki:
coarse_quantizer = faiss.IndexFlatL2 (d)
index = faiss.IndexIVFPQ (coarse_quantizer, d,
ncentroids, code_size, 8)
index.nprobe = 5
Elsewhere (e.g., this) you will see:
import faiss
d = 128 # Dimension (length) of vectors.
M = 32 # Number of connections that would be made for each new vertex during HNSW construction.
nlist = 10000 # Number of inverted lists (number of partitions or cells).
nsegment = 16 # Number of segments for product quantization (number of subquantizers).
nbit = 8 # Number of bits to encode each segment.
# Create the index.
coarse_quantizer = faiss.IndexHNSWFlat(d, M)
index = faiss.IndexIVFPQ(coarse_quantizer, d, nlist, nsegment, nbit)
Pay attention to the fourth parameter of IndexIVFPQ. In case of the first example, it is set equal to the code_size which is
M * nbits / 8 bytes (M subvectors taking up nbits each). In case of second example it is set equal to M. Which is correct?
It turns out the second example is the correct one and official documentation is wrong! Its not only there. I have seen the parameter
incorrectly referred to as code size (or the size of the compressed vector) at several other places in Faiss documentation.
Another example is found here:
PQM compresses the vectors using a product quantizer that outputs M-byte codes.
You can verify by inspecting the C++ source code that the fourth parameter is indeed M – the number of segments into which a vector should be divided.
When nbits = 8 which is the default and common setting, the code_size = M
so you will not catch the bug and the statement above is correct in a sense.
Maybe Faiss will correct their documentation at some point in future.
How not to create the IVFPQ index
Note that the coarse_quantizer input to IndexIVFPQ is of type IndexFlatL2. In particular it is not
a IndexIVF. An inspection of the
source code
reveals that the coarse_quantizer is being used to create a IndexIVF internally. See the C++ code copied from source below:
IndexIVFPQ::IndexIVFPQ(
Index* quantizer,
size_t d,
size_t nlist,
size_t M,
size_t nbits_per_idx,
MetricType metric)
: IndexIVF(quantizer, d, nlist, 0, metric), pq(d, M, nbits_per_idx)
Another thing to note: In the faiss Python package there is a class IndexIVF but its abstract. The C++ IndexIVF is not abstract.
If you try to create an index using the faiss index factory and pass it "IVFx" (e.g., "IVF100"), you will get error.
What about OPQ?
Maybe you have heard of Optimized Product Quantization and wondering what that is. Let’s revisit PQ. We have a vector, e.g.:
x = [0.77759472, 0.28507821, 0.58818803, 0.07639025, 0.90228833,
0.13530444, 0.35468633, 0.76250536, 0.83748744, 0.08922487]
and we split it into M = 5 subvectors in preparation of PQ:
>>> parts=[x[i*2:i*2+2] for i in range(5)]
>>> parts
[[0.77759472, 0.28507821], [0.58818803, 0.07639025], [0.90228833, 0.13530444], [0.35468633, 0.76250536], [0.83748744, 0.08922487]]
Why not split it some other way? E.g., what about splitting it as:
[[0.77759472, 0.58818803], [0.28507821, 0.07639025], [0.90228833, 0.35468633], [0.13530444, 0.76250536], [0.83748744, 0.08922487]]
do you see the difference? if not, read again. There could be many other ways to split and subdivide x into M parts.
What subdivision would minimize the quantization error between the original vectors and their quantized versions (stored as codewords)?
This is what OPQ or Optimized Product Quantization addresses. In general, we model this problem as applying a rotation matrix R
to x before quantizing it. OPQ finds the best R such that the quantization error between PQ(R(x)) and x is minimized when
aggregated over all values of x i.e., the entire dataset. There is a single rotation matrix R computed for entire dataset (not a
separate R for each data point – that would be too inefficient and probably overkill). Once the optimal R is computed in the training step,
it is applied to each vector before encoding it with PQ. That’s it for OPQ.
What about ScaNN?
ScaNN is a top-performing ANN search algorithm invented by Google.
It is same as IndexIVFPQ except that instead of computing the product quantization error as (within each subspace):